In this series we’re focusing on the integrative approach to preventing kidney stone formation.
Conventional approaches to kidney stones tend to focus on medications, surgical removal, and using ultrasonic waves to break up stone. It rarely approaches the root cause including risk factors to prevent stone formation.
We covered the impact of diet, the microbiome and gut health, and electrolyte imbalances on kidney stones in previous blogs. Here we will discuss the role genetics play in kidney stone formation.
The Genetics of Kidney Stones
There seems to be a familial link when it comes to the development of kidney stones. In fact, two thirds of patients with calcium-containing kidney stones have a relative with kidney stones. Recently, genome-wide association studies uncovered several genetic sequence variants (SNPs) that lead to increased risk of kidney stone development. Although we are still scratching the surface in understanding the contribution of genetic factors to stone formation, we do know that we can modulate these risks through environmental and dietary modifications.
Genetic Variations in Calcium Handling
In a previous blog, we discussed the role of the kidney filtration units (specifically the nephrons). The kidneys are responsible for filtering large volumes of blood daily. This function is crucial, and, the unique design of the nephrons make it able to adjust this filtrate and prevent dehydration. This intricate design also makes the kidneys crucial in the balance of water and many electrolytes.
Calcium is one of the electrolytes filtered and reabsorbed in the kidneys. Calcium sensing receptors (CaSR) are present in kidney cells and are essential for the reabsorption of calcium. These receptors increase and decrease the amount of calcium reabsorbed based on the calcium level in the blood. In other words, activating these receptors increases the amount of calcium lost in the urine.
Single nucleotide polymorphisms (SNPs) in CaSR have been found to alter the function of these receptors leading to increased urinary excretion of calcium. Of particular interest, SNPs in rs7652589 and rs1501899 were associated with kidney stones in patients with normal citrate excretion. SNPS in rs1801725, rs1042636 of the CaSR gene were also associated with kidney stones in various populations.
Another gene that has been associated with increased calcium in the urine is the gene coding for the protein Claudin-14, responsible for forming tight junctions. This protein helps connect adjacent cells to form a barrier which acts like a gate, separating blood from urine. Tight junctions are also responsible for ensuring that minerals don’t pass between the cells. SNPs in the genes that code Claudin-14 (CLDN14) alters the integrity of these “gates” and allows for calcium to “sneak” between cells into the urine, increasing the risk for kidney stone formation. Specifically, SNPs at the locations rs219778, and rs219780 of the CLDN14 gene were significantly associated with kidney stones.
Genetic Variations in Vitamin D Receptors (VDR)
Vitamin D plays a crucial role in calcium balance. Studies have shown that vitamin D increases the absorption of calcium from the gut and also increases calcium excretion in the urine. Vitamin D receptors are essential for vitamin D to exert its action on calcium balance.
Some mutations or SNPs in the VDR are associated with increased absorption and excretion of calcium, significantly increasing the risk of kidney stones. It is worth mentioning here that there is some controversy about the link between vitamin D supplementation and kidney stone formation. Vitamin D deficiency appears to be common among kidney stone formers. This is likely because low vitamin D causes calcium loss from the bone in order to maintain normal calcium range in the blood for cardiovascular protection. Even though vitamin D3 supplementation may increase calcium excretion in the urine, it has not been conclusively found to increase the risk of kidney stone formation.
Therefore, genetic assessment may be a key to identify patients who are at risk of kidney stone formation from taking vitamin D supplements.
Genetic Variations in the Handling of Other Minerals
The kidneys are crucial for balancing many minerals in our bodies such as magnesium, phosphate, oxalate and others. Genetic mutation or SNPs affecting the genes that code for the channels or receptors that regulate these minerals can also impact the risk of kidney stones. Some SNPs on the other hand can be protective against kidney stones such as SNPs in the UMOD gene. Discussion of this long list of SNPs requires details beyond the scope of this blog, but we summarized most of the genes that have been associated with kidney stones in the table below.
| Gene symbol | Gene name | Phenotype |
| ADCT10 | Adenylate cyclase 10 | Increased calcium excretion |
| AGXT | Alanine-glyoxylate aminotransferase | Increased oxalate excretion |
| CA2 | Carbonic anhydrase II | Osteoporosis + decreased acid excretion |
| CASR | Calcium-sensing receptor | Increased calcium excretion |
| CLCN5 | Chloride channel, voltage-sensitive 5 | Dent disease |
| CLCNKB | Chloride channel, voltage-sensitive Kb | Bartter Syndrome, type 3 |
| CLDN14 | Claudin 14 | Increased calcium excretion |
| CLDN16 | Claudin 16 | Increased calcium and magnesium excretion |
| CLDN19 | Claudin 19 | Increased calcium and magnesium excretion |
| CYP24A1 | Cytochrome P450 | Decreased breakdown of vitamin D3 |
| GRHPR | Glyoxylate reductase | Increased oxalate excretion |
| HOGA1 | 4-Hydroxy-2-oxoglutarate aldolase 1 | Increased oxalate excretion |
| HPRT1 | Hypoxanthine phosphoribosyltransferase 1 | Increased uric acid excretion |
| SLC12A1 | Solute carrier family 12, member 1 | Bartter syndrome, type 1 |
| SLC26A1 | Solute carrier family 26, member 1 | Calcium oxalate kidney stones |
| SLC22A12 | Solute carrier family 22, member 12 | Decrease uric acid excretion |
| SLC2A9 | Solute carrier family 2, member 9 | Decreased uric acid excretion |
| SLC34A1 | Solute carrier family 34, member 1 | Calcium phosphate kidney stones |
| SLC34A3 | Solute carrier family 34, member 3 | Calcium phosphate kidney stones |
| SCL3A1 | Solute carrier family 3, member 1 | Increased Cystine excretion |
| SLC4A1 | Solute carrier family 4, member 1 | Decrease acid excretion (dRTA) |
| SLC7A9 | Solute carrier family 7, member 9 | Increased Cystine excretion |
| SLC9A3R1 | Solute carrier family 9, subfamily A, member 3, regulator 1 | Calcium phosphate kidney stones |
| UMOD | Uromodulin (most common urine protein) | Protective against kidney stones |
| VDR | Vitamin D (1,25-dihydroxy D3) receptor | Increased calcium excretion |
| XDH | Xanthine dehydrogenase | Increased xanthine excretion |
The Bottom Line
There are many factors that impact the risk of kidney stone development. Although there are pure genetic diseases that are associated with kidney stones, often the increased risk is subtle or offset by other factors. Increased risk, when combined with other factors including nutrient depletion, dysbiosis, electrolyte imbalances and dehydration, may lead to the development of kidney stones in some. Assessing the genetic profile of kidney stone patients can help identify the root cause of the disease to tailor appropriate, personalized management. Practitioners working with individuals to prevent kidney stone formation should formulate a comprehensive and individualized intervention that modifies all relevant components in their integrative approach.
The post Genetics of Kidney Stones appeared first on Integrative Kidney.]]>By Majd Isreb, MD, FACP, FASN, IFMCP
Exposure to Arsenic has been associated with increased risk for cancer and it is also linked to kidney disease. In this blog we discuss the sources of arsenic exposure, how our body handle arsenic, its effects on the kidneys, and the genetic and epigenetics of arsenic.
Sources of exposure
Drinking water appears to be the greatest source of arsenic exposure worldwide. In addition, exposure from ingested foods comes from food crops grown in arsenic-contaminated soil or irrigated with arsenic-contaminated water. Even if the water you drink is filtered and does not have arsenic, it is possible that the food that you eat is contaminated with arsenic, particularly rice. Arsenic is also found in high concentrations in cigarettes.
Even if the water you drink is filtered and does not have arsenic, it is possible that the food that you eat is contaminated with arsenic, particularly rice
Share on X
Found naturally in the environment, arsenic is mobile and cannot be destroyed. When arsenic compounds interact with oxygen or other molecules or bacteria it changes into different forms that can dissolve in water. Cosmetics and personal care products are another often overlooked major source of exposure.
How our body detoxifies arsenic
After ingestion or exposure, arsenic is metabolized and detoxified by a process called methylation. An enzyme called arsenic methyltransferase (AS3MT) adds a methyl groups to arsenic compounds to help neutralize it and eliminate it. This enzyme has been detected in human liver, kidney, bladder, heart, lung, testes and adrenal glands. Glutathione, known as the master antioxidant also plays a key role in arsenic detoxification.
The various arsenic forms bind to glutathione. The compound arsenic-glutathione can easily exit the cells and circulate in the blood. The glutathione-bound arsenic compound can be excreted by the kidneys or in the bile. The kidneys are the major site of elimination of arsenic and they are, therefore, highly exposed to it. Arsenic can also accumulate in the kidneys. Many transporters have been implicated in arsenic excretion in the tubules portion of the filtering units of the kidneys (the nephron). Inhibition or genetic changes in these transporters can lead to enhanced arsenic toxicity.
The kidneys are the major site of elimination of arsenic and they are, therefore, highly exposed to it. Arsenic can also accumulate in the kidneys.
Share on X
Interestingly, the water channel, aquaporin 3, has been found to increase the uptake of arsenic in the kidneys. These channels play a crucial role in water handling by the kidneys. They help our body preserve water. Increasing water intake decrease the incorporation of these channels to the cells of the tubules in the kidneys. Therefore, increasing filtered water intake can help decreasing the kidney exposure to arsenic.
Finally, a transporter called multidrug resistance transport protein 2 (MRP2) transport arsenic-glutathione complexes and excrete them out of kidney cells into the urine. Studies show that the increase in the production of glutathione and MRP-2 increase the body’s ability to detoxify arsenic. Selenium is important for this process and selenium deficiency can enhance arsenic toxicity.
Selenium and zinc deficiencies can enhance arsenic toxicity.
Share on X
Arsenic effects on the kidneys
Acute exposure of the kidneys to toxic levels of arsenic leads to inflammation in the tubules of the kidneys. This can lead to protein loss in the urine. It can also lead to elevated calcium levels in the blood. Arsenic exposure activates cell growth can lead to cancer. Arsenic can also lead to oxidative stress and cellular injury. Arjunolic acid (an anti-oxidant which is present in fruit specially guava) was found to decrease the toxicity of the arsenic on the kidneys. Zinc is also protective. Chronic exposure can lead to high blood pressure, protein loss in the urine, and chronic kidney disease.
Download Your Copy!
Genetics and Genomics Impact on Arsenic Toxicity
You can now see that various processes are involved in the handling of arsenic by the body from detoxification to transport to elimination. Genetic variants of any of these steps can either be protective or can increase arsenic toxicity. For example, a single nucleotide polymorphism (SNP), or genetic mutation, that increases glutathione production can be protective since glutathione helps to eliminate arsenic safely. However, a SNP in the gene that codes for MRP2 may potentially decrease arsenic excretion leading to increased toxicity.
Arsenic accumulation and toxicity can lead to oxidative DNA damage. It also inhibits DNA repair. This explains the increased risk for cancer after exposure to arsenic. Chronic exposure to even small concentrations of arsenic can have synergistic effect on other factors that can cause cancer such as UV light and smoking.
Arsenic exposure can cause indirect DNA damage through processes of DNA methylation and histone acetylation. These altered epigenetic expressions have been linked to kidney inflammation and damage. More recently, it was noted that long term exposure to low level of arsenic (below what is considered safe by the EPA guidelines) were associated with epigenetic changes that caused kidney scarring.
it was noted that long term exposure to low level of arsenic (below what is considered safe by the EPA guidelines) were associated with epigenetic changes that caused kidney scarring.
Share on X
The Bottom Line
Most arsenic exposure is due to drinking and showering with unfiltered water. Arsenic exposure leads to extensive cellular damage which increases the risk for cancer and kidney disease. Nutritional deficiencies can enhance its toxicity, but you can take steps to reduce your risks.
- Choose a home filtration device that effectively removes heavy metals, including arsenic. Not all consumer filters will remove arsenic, consider upgrading to Clearly filtered, Multipure and Aquasauna water filter systems.
- Always choose organic produce whenever possible, However, when it comes to rice and teas – organic isn’t enough. Look for products that screen for arsenic levels.
- Antioxidants are protective against arsenic damage, so a diet high in antioxidants like citrus fruit, green tea, blueberries, and dark chocolate will be naturally protective. Enhance detoxification by eating dark leafy greens, cruciferous veggies (like broccoli, kale, and cabbage), and sulforaphane-rich foods like garlic and onion contain high concentrations of nutrients that increase our detoxification capacity.
- Load up on fiber and mineral-rich root veggies, which help trap and eliminate toxins through your digestive system.
- There are several lifestyle modifications that can support improved detoxification, including Epsom-salt baths, sauna, lymphatic massage, sweat-inducing workouts, and dry brushing.
- If like many Americans you’re not having consistent daily bowel movements, you’re not adequately detoxifying and should consider working with a healthcare provider to address your digestion and motility.
Foot Note:
There are different forms of arsenic compounds. There are organic and inorganic forms with different oxidation states. The inorganic forms are easier to absorb by the gut and, therefore, are more hazardous. The most commonly ingested inorganic compounds are the trivalent and the pentavalent forms (depending on the level of oxidation). The trivalent forms appear to be the most toxic.
The post Arsenic and Kidney Health appeared first on Integrative Kidney.]]>2.2 million adverse drug reactions occur in the United States annually, and medication efficacy rates vary considerably. Pharmacogenomics offers the solution for safe and highly personalized drug therapy
Share on X
By Lara Zakaria, PharmD, CNS, CDN, IFMCP
Variability in response to pharmaceutical therapy is due to multiple, complex individual variations in basic physiological differences like age, sex, and weight, as well as metabolism, absorption, organ function, and disease state. These variables mean patients require monitoring, dose adjustments, and sometimes alternative therapy (drug switch).
This is why medications come in various doses, formulations, and options exist within a drug class. Prescribers follow “best practice guidelines” that allow them to choose an alternative when one drug therapy fails or is not tolerated due to side effects or ADR. But even with these well-researched guidelines, the choice and dosage of a medication can still be a guess.
Wouldn’t it be a relief to be able to predict in advance how someone might respond to medication, avoiding guessing, saving time, and improving patient outcomes?
Thanks to advances in the field of genetics called pharmacogenomics (PGx), clinicians are starting to use genetic information to personalize drug therapy. Though this has a broad range of benefits, our focus here will be on how this impacts those who have kidney disease (CKD) or kidney transplants.
Prescribers follow “best practice guidelines” … But even with these well-researched guidelines, the choice and dosage of a medication can still be a guess. Wouldn’t it be a relief to be able to predict in advance how someone might respond…
Share on X
Defining Pharmacogenomics
DNA is made up of a sequence of four nucleotides, (adenine, cytosine, guanine, and thymine). The genetic sequence serves as a template that our cells use to make thousands of proteins which are essential for carrying out all biological functions that maintain life. This includes the enzymes, receptors, and channels that are involved in the absorption, transport, and metabolism of drugs.
Alterations in the sequence, like single nucleotide polymorphisms (SNPs), or errors in the original code, may affect the efficiency of protein production. In turn, this may alter the efficacy or toxicity of a medication for an individual with that SNP.
When scientists began exploring pharmacogenetics in the 1950s, clinical research focused on connecting single genetic variations with a predictable outcome in drug therapy. In the last decade and a half, it has led to an evolution of pharmacogenomics, exploring the complex interplay of genetic and environmental influences on individual genetic expression.
Pharmacogenomics explores complex interplay of genetic and environmental influences on individual genetic expression
Share on X
Aside from being a relatively new area of focus, this nuanced difference is what makes “omics” research more challenging to translate into clinical application. Even though we’re only at the tip of the proverbial iceberg of this revolutionary approach to individualized medicine, useful clinical implications are already emerging, in particular when it comes to medications often used in kidney patients.
Last 20 years yielded a rapid influx of genomic data, with over 20,000 new #pharmacogenomics citations in PubMed, 3500+ gene-drug variant associations reported, and ~200 medications with FDA approved Pgx labeling
Share on X
Pharmacogenomics of kidney disease management
To date, there are approximately 300+ medications with FDA-approved PGx labeling recommendations. Of these, a handful is common in kidney disease patients. These categories include*:
- Cardiovascular Disease (CVD) – including clopidogrel (CYP2C19), simvastatin (SLOCO1B1) among other HMG-COA inhibitors, also known as “statins,” and warfarin (multiple, including CYP2C9, CYP4F2, VOKORC1, and potentially APOE, ABCB1, and UGT1A1)
- Transplant medications – including azathioprine (TPMT), tacrolimus (CYP3A4/3A5), and the antifungal agent voriconazole (CYP2C19)
- Hyperuricemia (elevated circulating uric acid) – allopurinol (HLA-B)
- Diabetes, with metformin being the most well-known (SNP rs11212617).
| Therapeutic Category | Drug | Genetic SNP |
| Cardiovascular Disease (CVD) | Clopidogrel | CYP2C19 |
| HMG-CoA inhibitors (statins) | SLOCO1B1 | |
| Warfarin | CYP2C9, CYP4F2, VOKORC1, and potentially APOE, ABCB1, and UGT1A1 | |
| Transplant medications | Azathioprine | TPMT |
| Tacrolimus | CYP3A4/3A5 | |
| voriconazole (antifungal agent) | CYP2C19 | |
| Hyperuricemia (elevated circulating urea) | Allopurinol | HLA-B*58:01 |
| Diabetes | Metformin | SNP rs11212617 |
* This is a list narrowed down limited to the strength of available evidence and relevance to KD by Adams et al.
The need for individualized therapy in kidney patients
Even when dosing is in line with best practice guidelines, they may still be sub-optimal or nephrotoxic (toxic to the kidney). Either situation for a transplant or KD patient may be life-threatening. Knowing in advance how a patient will respond to therapy means avoiding the cycle of “guess, assess and adjust,” saving valuable time.
Let’s use the medication tacrolimus, as an example. Tacrolimus is a very widely used and preferred immunosuppressant medication used in patients after a kidney transplant. Monitoring is very important because there’s a small margin of error for dosing. Due to this narrow therapeutic window, a little too much of the drug can quickly result in toxicity, while slightly under-dosing will be ineffective and potentially cause transplant rejection. Either situation can be life-threatening for a transplant patient.
Tacrolimus is metabolized by enzymes (part of the cytochrome P450 class) referred to as CYP3A4 and CYP3A5. It’s understood that individuals considered “hyper-metabolizers” – enzymes work faster than average to metabolize (break down) the drug – require higher dosing to achieve therapeutic efficacy. Meanwhile, slow metabolizers, require smaller doses to maintain safe and effective therapeutic levels. Because of the narrow therapeutic window, there is a small margin for accuracy, making these variations in metabolic functions particularly relevant to safety and outcome.
It’s interesting to note that there are various factors that influence CYP3A4/3A5 rate of metabolism. Use of other common medications like acetaminophen, clarithromycin, SSRI antidepressants (like fluoxetine and sertraline), and even caffeine, cannabis, and phytonutrient compounds naturally found in foods and herbs can also serve to up or downregulate the function of CYP3A4/3A5– further confounding or contributing to successfully achieving the narrow therapeutic window.
Another interesting example is the case of the transporter gene SLCO1B1 and its effect on statin therapy. This anti-cholesterol medication class (which includes simvastatin, rosuvastatin, pravastatin, etc) is well known for an ADR which causes muscle breakdown: rhabdomyolysis. Risk increases with increased blood concentration or dosage. Furthermore, muscle breakdown can add stress to the kidney.
SLOCO1B1 is responsible for transporting the drug out of the blood and into the liver for detoxification. In individuals with genetic variations that reduce the function of the transporter, accumulations are more likely to occur, and rhabdomyolysis is more likely even at lower drug dosage than typically expected. Combine this with environmental factors or combination drug therapy that may further slow detoxification, liver function, or drug clearance, and you can start to see how multiple factors may contribute to presentations of this ADR.
Should you get genetic testing?
There is a multitude of consumer tests offering direct-to-consumer genomic and health data like 23andMe, among others. The health reports are limited, and may omit some crucial lifestyle and epigenetic factors, but can be a good starting point. Raw SNP data can be obtained, but interpretation requires a strong understanding of clinical significance so working directly with a healthcare professional with appropriate training is strongly recommended.
There are also professional tests and reports available through your health care provider like IntellexxDNA and MyDNA. Your PGx-literate provider can help you choose the appropriate test, and most importantly guide you in recommendations once you have your results in hand.
It’s essential to remember that the field of PGx is relatively young and complex. When we talk about genetics, we cannot ignore the significance of epigenetics. In a previous blog, we discussed how dietary and lifestyle modifications can affect expressions of these genetic traits.
Another related emerging field is nutrigenomics– a cousin to PGx – focusing on the effect of nutrients on genetic expression. When interpreting emerging findings in omics, clinicians must take into consideration the effect of factors like nutrition, herbal use, as well as environmental toxins, and medication on epigenetics expression. The individualized and integrative approach to managing kidney disease will rely on the nephrologists, pharmacists, and clinical nutritionists of the not-so-distant future working together to layer those factors to maximize therapeutic benefit and reduce harm to patients.
The post Pharmacogenomics: Advances in Individualized Treatment in Kidney Disease appeared first on Integrative Kidney.]]>What is IgA?
Immunoglobulins are a large protein structure that is part of the adaptive immune system. There are various types, categorized by a letter and sometimes a number to identify the subclass, for example IgA1and IgA2. For the context of this discussion, we will the focus immunoglobulin A (IgA) without distinction to subclass.
IgA is found in the blood serum, lining of the respiratory (lungs) and digestive (gut) tracts. It is also found in saliva, tears, and breastmilk. Also called an antibody, the body makes IgA and other types of antibodies to help fight off invading pathogens (i.e. bacteria, viruses) and prevent disease. It is one of the first lines of defense against invading organisms from the environment, diet, and toxins.
What is IgA nephropathy?
IgA nephropathy is an autoimmune kidney disease. A trigger causes the immune system to produce abnormal IgA proteins that causes the immune defense to damage the kidney (nephrons). Abnormal IgA immune complex structures are formed, they circulate and filter through the kidney where they are “captured”. Unfortunately, these structures build-up in the kidneys, resulting in inflammation which damages kidney tissues.
What triggers an IgA immune response?
Though there’s some evidence that points to certain heavy metals (like cadmium) as a source of toxicity, the strongest evidence in the literature points to triggers can be classified into the following categories:
- Food sensitivities/allergies (i.e. gluten, soy, lectins/lipopolysaccharides)
- Microbial imbalance (known as dysbiosis) in the intestinal tract
Several studies have demonstrated an association between intake of gluten and lectins. One study demonstrated that patients with IgA nephropathy who followed a gluten-free diet for 6 months had a reduction in IgA levels. Studies on Mediterranean people eating a diet rich in gluten-containing foods published as far back as 1986 have suggested a dietary component to kidney disease. Soy is another example of a food antigen that has been associated with IgA nephropathy. However, heavy use of pesticides (glyphosate) in our food sources (especially gluten and soy) may play a role in food-related nephropathy.
Heavy use of pesticides (glyphosate) in our food sources (especially gluten and soy) may play a role in food-related nephropathy
Share on X
In susceptible individuals, the intake of a dietary trigger leads to intestinal permeability (IP), also known as leaky gut. In fact, research dating back decades has supported a connection between the gut and IgA nephropathy, called the gut-kidney connection. One of the proposed mechanisms, is the development of microbial imbalance (dysbiosis) which leads to bacteria and food proteins “leaking” through the gut barrier, triggering IgA immune response damaging the kidney.
The role of dysbiosis
In the case of IP, the compromised lining of the gut (mucosa) allows abnormal microbes, endotoxins (toxins produced by microbes), or food particles and metabolites to enter the bloodstream.
Furthermore, studies have shown that patients with progressive IgA nephropathy tend to have low number of naturally occurring healthy bacteria in the gut (likeBifidobacterium species) and a high amount of unwanted, or pathogenic, bacteria (likeSterptococceae). Our gut lining cells have complex mechanisms to detect the type of microbes that are present in our gut. There is a consistent link between these mechanisms and the production of an abnormal IgA.
The genetic link to an abnormal IgA response?
Genetics likely also play a role in the risk of developing kidney disease. Researchers have identified genetic variances (also referred to as single nucleotide polymorphisms, or SNPs) linked to IgA nephropathy in a multinational study.
Most of the genes involved in regulating the response and immunity against gut microbes. Interestingly, many of these SNPs associated with IgA nephropathy risk are related to inflammatory bowel diseases (IBD) or with the maintenance of the intestinal barrier. These SNPs lead to abnormal trafficking of IgA and the formation of an abnormal IgA that triggers the body’s immune response.
The autoimmune response, putting it all together
Recall, the presence of abnormal IgA in the blood leads to the formation of protein complexes in response to neutralize the threat. Unfortunately, these proteins end up forming structures that are difficult for our body to eliminate. These complex structures with abnormal IgA end up depositing in the kidney, leading to local inflammation and loss of kidney function.
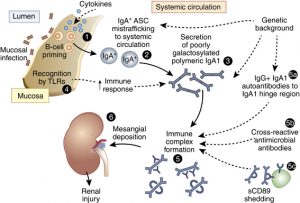
Now it should be clearer that IgA nephropathy is a classic example of how a combination of genetic, environmental factors, dietary and gut integrity interact to cause disease.
IgA nephropathy is a classic example of how a combination of genetic, environmental factors, dietary and gut integrity interact to cause disease
Share on X
Conventional medicine approach
The conventional approaches have failed to find a completely effective treatment for IgA nephropathy. Approaches include conservative control of blood pressure and blockage of the angiotensin-renin pathway, reduction of protein in the urine, and use of immunosuppressive therapy (including steroids and other medications), and fish oil.
More recently, a special kind of steroid that targets the terminal part of the gut (ileum and cecum) is being studied. The hope is that these special steroids will focus on one aspect of the process that leads to the formation of abnormal IgA which is decreasing the immune response in the gut.
Integrative medicine approach
A more integrative approach to IgA nephropathy would personalize treatment to take into account the impact of IP:
- Identifying the and eliminating the dietary or environmental triggers that contribute to leaky gut
- Assess and address dysbiosis
- Capitalize on epigenetic modifications to reduce the expression of SNPs contributing to IgA upregulation.
In combination with evidence-based conventional medicine, this approach targets various mechanisms and complications of the disease and improving outcomes.
The post IgA nephropathy: A target for Integrative approach appeared first on Integrative Kidney.]]>





